Introduction
Salmonella spp. is the most relevant foodborne pathogen globally. It is widely distributed in nature and it is able to survive in a wide range of environments. Human infections are associated not only with animal food sources, but also with pets, reptiles, fruits, vegetables, and other humans. However, poultry and poultry products are considered as one of the most important sources of human Salmonella infections (Batz et al., 2012).
More than 2,500 Salmonella serovars are currently identified, and about 100 are recognized relevant to public health (WHO, 2013; CDC, 2015). The ability of different Salmonella serotypes to survive and thrive in different host environments involves several interconnected factors, including differences in host environment, in the immune system response to the different serotypes, the presence of commensal organisms, and the genetics of the pathogen itself (Foley et al., 2013).
Prevention, control, and monitoring programs are powerful tools to reduce the presence of Salmonella in the poultry production chain (Santos et al., 2013); however, the knowledge on the contribution of different serotypes and of contamination sources is essential to determine food safety intervention priorities and to implement appropriate control measures (Campos et al., 2012). Understanding the mechanisms of Salmonella pathogenesis is important for the development of intervention strategies aiming at limiting its prevalence and spread, as well as of novel drugs and vaccines that could provide better treatment and/or prevention of salmonellosis in animals and humans (Foley et al., 2013).
Genotyping is important tool in epidemiological studies and for tracing foodborne pathogens. Pulsed Field Gel Electrophoresis (PFGE) method is considered the gold standard in genotyping (Ribot, 2006). However, laboratory data must match epidemiological and environmental information because PFGE results alone cannot establish an epidemiological connection among isolates, and therefore, all available information must be considered when interpreting genetic subtyping results (Barret et al., 2006).
The objectives of this study were: 1) to compare the effectiveness of the same Salmonella prevention and control program applied in two different broiler processing plants belonging to the same company and located in the states of Mato Grosso do Sul (plant A) and Santa Catarina (plant B), Brazil, considering the food safety of frozen chicken breast; and 2) to identify the Salmonella strains isolated from live broilers, carcasses before and after chilling, and frozen chicken breasts.
Materials and methods
In total, 1800 carcasses from 20 broiler flocks slaughtered in two processing plants belonging to a single company (plants A and B, located in the states of Mato Grosso do Sul and Santa Catarina respectively), between 2012 and 2013, were evaluated for Salmonella spp. The company applies identical programs for internal quality assurance in both plants, which have high hygiene standards. In addition, the company applies the same program for the prevention, control, and monitoring Salmonella in its processing plants and broiler farms. The efficacy of the Salmonella prevention and control programs were evaluated considering a maximum of 10% Salmonella-positive samples per batch of frozen chicken breasts. Sampling was performed according to Van Schothorst et al. (2009).
The selected flocks were monitored pre-harvest by isolation of Salmonella from drag swabs, and the results showed that 50% were positive and 50% negative. The samples were collected 15 days before slaughter, placed in sterile bags, and submitted to the in-house animal health laboratory on ice. Samples were processed within 24 h of collection, and Salmonella strains were identified according to the official methods of the Brazilian Ministry of Agriculture (Brasil, 2009).
At the processing plant, 10 cloacal swabs (each swab was used in 5 birds, totaling 50 birds) were collected from each flock at the reception platform and placed into sterile tubs with 25 mL of BPW at 1%. After slaughter,25 carcasses per flock were collected before and 25 after the chiller, placed individually into a sterile plastic bag, and transported to the laboratory on ice and processed within 24 hours. Thirty frozen chicken breasts per flock were collected after 30 days of storage at -18°C, and submitted to the laboratory. The carcasses and chicken breast were cut, weighed, and homogenized in a stomacher under sterile conditions in a laminar flow chamber.
Carcass and breast samples weighing 25 g were placed in 250 mL 1% BPW.
Cloacal swabs, and carcass and breast samples were pre-enriched in1%BPW at 37oC for 18-24 h. Cloacal samples were then regrown in brain heart infusion broth (BHI; Difco, Detroit, MI, USA) at 37oC for further 4hours. Salmonella spp. was detected following the protocol of the automated PCR BAX System (DUPONT QUALICOM, USA). Salmonella colonies were enumerated in 7.5 mL pre-enriched samples by the miniaturized most probable number (mMPN) technique, according to ISO6579-2 Part 2.
Positive samples were confirmed by bacteriological isolation on Rambach agar (Beacton Dickinson, USA) and incubated at 37oC for 24 h. Suspected colonies were confirmed by biochemical and seroagglutination assays using polyvalent antiserum.
The strains isolated in mNMP (116 samples), the drag swabs from the evaluated broiler flocks (10), and three strains isolated in 2012 from drag swabs collected on farms supplying plant A were purified in1% BPW, transferred to nutrient agar tubes, and submitted to the Enterobacteria National Reference Laboratory, Instituto Oswaldo Cruz (IOC), where complete antigen characterization, serotype identification, and genotyping by pulsed-field gel electrophoresis (PFGE) were performed.
Genotyping protocol was applied according to the Center of Disease Control of the PulseNet Network (CDC, 2013). The isolates were genotyped by DNA macrorestriction analysis, using 40 U of the enzyme XbaI (New England Biolabs, Beverly, MA, USA) followed by PFGE, as previously described (Ribot et al., 2006). Salmonella Braendrup was used as size standard. Restriction fragments were electrophoresed in certificated 1.2% PFGE agarosegels (Bio-Rad, Hercules, CA, USA) in tris-borate buffer (TBE; trisborate 0.045 M, EDTA 0.001M) at 140C, using the CHEF DR III system (Bio-Rad), with an initial switch time of 2.2s and a final switch of 63.8s at 6V/s for 18 h. Gels were stained in ethidium bromide (1μlg/ mL) and visualized under UV light. Images were captured using a digital camera, and macrorestriction patterns were compared using BioNumerics 4.0 software (version 6.1) (AppliedMaths, Sint-Martens- Latem, Belgium). Similarity was calculated by the Dice coefficient with 1.7% tolerance. A dendrogram was generated by cluster analysis using the unweighted pair of group method with arithmetic mean (UPGMA). Strains sharing the same number and position of DNA macrorestriction fragments were considered to belong to the same genotype.
Results and discussion
Out of 1,800 cloacal swab, carcass before and after chilling, and frozen breast samples of the 20 broiler flocks evaluated, 278 samples were positive for Salmonella spp by A-PCR and 118 samples were enumerated by mNMP.
Salmonella spp was detected by A-PCR in 225/900 (25%) samples from plant A and in 53/900 (5.88%) from plant B, and 116/900 (12.9%) were enumerated by mNMP in plant A samples and 2/900 (0,22%) in plant B samples. Table 1 shows the number of samples positive for Salmonella spp and enumerated at different collection steps.
Table 1 – Frequency of Salmonella-positive samples detected by automated PCR (A-PCR) and frequency of enumerated samples by miniaturized most probable number (mNMP) per flock obtained in two poultry processing plants located in the states of Mato Grosso do Sul (plant A) and Santa Catarina (plant B), Brazil.
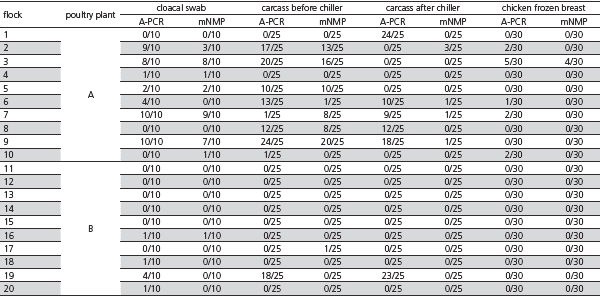
The results show that the Salmonella prevention and control program was 90% effective in Plant A and 100% effective in Plant B, considering the number of Salmonella-positive frozen chicken breast samples. The number of 30 samples and the acceptance limit of 10% Salmonella-positive samples per frozen chicken breast batch are established by International Commission on Microbiological Specification for Foods (ICMSF), and indicates that a proportion of 0.18% contaminated carcasses is acceptable at 95% probability level (Van Schothorst, 2009). In batch 3 of frozen chicken breasts processed in plant A, the number of Salmonella-positive samples 5/30 as detected by A-PCR and 4/30 as enumerated by mNMP. The combination of qualitative and quantitative results make clear the importance of the level of contamination to food safety of chicken breast (Straver et al., 2007).
In total, 128 strains were serotyped: 10 from drag swabs, 31 from cloacal swabs, 83 from carcasses(77 before and 6 after chilling), and 4 from frozen chicken breasts. Nine nontyphoidal Salmonella serotypes were identified: Salmonella Minnesota; Salmonella Newport; Salmonella Senftenberg; Salmonella Agona; Salmonella Anatum; Salmonella Mbandaka; Salmonella entérica susp. enterica O:4,5; Salmonella enterica subsp. enterica O:6,7:-:e,n,z15; and Salmonella enterica subsp. enterica rough.
After serotyping analysis, 30 strains selected at different processing steps and drag swabs, and three Salmonella Minnesota strains isolated from plant A in 2012 were genotyped by PFGE, which identified 5 non typhoidal Salmonella: Salmonella Minnesota, Salmonella Newport, Salmonella Senftenberg, Salmonella Anatum, and Salmonella Typhimurium.
Salmonella detection frequency and enumeration were higher in plant A samples, and therefore, 94.4% of serotyped samples originated from Mato Grosso do Sul. In this plant, the most frequently isolated strain was Salmonella Minnesota, with 111/121 (91.3%), followed by Salmonella Newport with 10/121 (8.77%). Two strains were serotyped as Salmonella enterica susp. enterica (O:6,7:-:e,n,z15),and Salmonella enterica susp. enterica rought however in the PFGE were identified as Salmonella Minnesota.
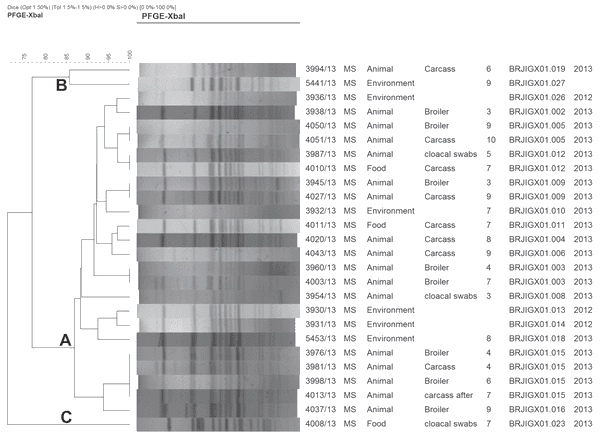
Salmonella Minnesota strains were differentiated by PFGE in 19 pulsotypes (Figure 1) distributed in three clusters (A, B and C), as shown in Figure 1.
Figure 1 – XbaI PFGE of Salmonella Minnesota isolates from drag swabs, cloacal swabs, carcasses and breast obtained in plant A, state of Mato Grosso do Sul, Brazil.
Cluster A was predominant and endemic in the region. The comparison of the strains isolated in our study with the three strains isolated from drag swabs obtained in poultry farms supplying plant A in 2012 showed a temporal and spatial relation among the pulsotypes. The older strains belonged to cluster A, indicating these clones are very stable and did not significantly changed over time.
The low representativeness/frequency of B and C clusters may be due to a novel transmission route or changes in the processing procedures, allowing the development new clusters or genetical changes. We do not have epidemiological data to interpret this finding.
Salmonella Minnesota represented more than 90% of all isolates, demonstrating it was dominant and endemic in all evaluated steps of processing. Salmonella Minnesota has been increasingly isolated in the Brazilian poultry chain since 2009, particularly in the Midwestern region (Santos, 2013; Cardoso & Tessari, 2015). Voss-Rech et al. (2015) reported a high frequency of Salmonella Minnesota isolation in broiler farms located in Mato Grosso do Sul, and suggested that this serotype is predominant and widely distributed in that region. Our findings support this hypothesis.
The environmental and ecologic conditions of the Brazilian Midwest, as well as the long distances between the farms and the processing plants represent an additional challenge to implement effective Salmonella prevention and control programs. During the crop planting season, in particular, the presence of reptiles and wild birds around processing plants and broiler farms increases due to the presence of feed. The Ecology and Animal Epidemiology session of the 13thInternational Symposium Salmonella and Salmonellosis, held in France in 2013, concluded that wild birds and reptiles may be considered as new epidemiological routes of Salmonella (International Symposium Salmonella and Salmonellosis, 2103).
Pickler et al. 2012 evaluated the effectiveness of the supplementation of broilers with organic acids in the feed and drinking water during the pre-harvest period for the prevention of Salmonella Minnesota and Salmonella Enteritidis contamination, and showed that organic acids were not effective against Salmonella Minnesota. The use of organic acid during the preharvest period is recommended by international guidelines (CODEX, 2011; OIE, 2015), and it is adopted by many Brazilian poultry companies. However, in the specific case of Salmonella Minnesota endemic region, it may result in the selection of resistant strains and the dominance of this serovar.
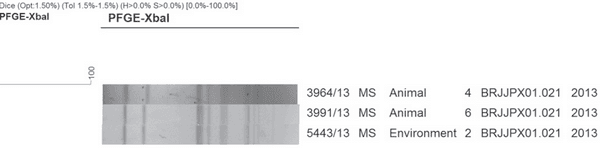
Three strains of Salmonella Newport were genotyped and identified as a single clone, with 100% of genetic similarity (Figure 2). This clone belongs the cluster isolated from broiler breeders in the Brazilian Midwest since 2009, and it was identified in human salmonellosis outbreaks in Minas Gerais and Rio Grande do Sul states in 2006.
Figure 2 – XbaIPFGE of Salmonella Newport isolates from drag swab, cloacal swab, carcasses from plant A, Mato Grosso do Sul, Brazil.
In plant B, low frequency of detection of Salmonella by A-PCR and enumeration by mNMP were determined in the samples obtained in all evaluated steps. Only seven strains were isolated: five from drag swabs, one from cloacal swabs, and one from carcasses. None Salmonella was isolated from frozen breast samples in plant B.
The phenotypic identification by serotyping of the four strains genotyped by PFGE diverged from the genotypic results. The three strains serotyped as Salmonella Agona, Salmonella Mbandaka, and Salmonella enterica susp. enterica rought were genotyped as Salmonella Senftenberg, and one SalmonellaTyphimurium pulsotype was serotyped as Salmonella enterica susp. enterica O:4,5.It should be emphasized that genotyping methods were developed to overcome the disadvantages of traditional phenotypic methods, particularly the influence of environmental factors (Vernile et al. 2009).
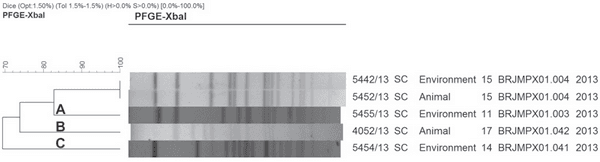
Salmonella Senftenberg was isolated in 80% (4/5) of the drag swabs from Salmonella-positive flocks in pre-harvest monitoring, and in a single cloacal swab sample. Salmonella Senftenberg was differentiated by PFGE in four pulsotypes distributed in three clusters (A, B, and C) as shown in Figure 3. Cluster A presented three samples with two pulsotypes.
Figure 3 – XbaIPFGE of Salmonella Senftenberg isolates from drag swabs and cloacal swabs obtained from plant B, Santa Catarina, Brazil.
The strain serotyped as Salmonella Agona was identified by PFGE as a Salmonella Senftenberg pulsotype. Salmonella Agona and Salmonella Senftenberg present three serologic factors in the scheme of identification: two of them are similar and, in some cases, there is a cross reaction (personal communication). The epidemiological context is relevant to evaluate the results.
The strain isolated from drag swabs serotyped as Salmonella enterica subs. enterica O:4,5 lost its flagellum, and therefore its complete identification was not possible; however, it was identified as a Salmonella Thyphimurium pulse type by genotyping. Salmonella Anatum was serotyped and genotyped in a single sample of carcass before chiller. No related relevant epidemiological data were identified (Figure 4).
Figure 4 – XbaIPFGE of Salmonella Typhimurium and Salmonella Anatum isolates from drag swab and carcasses before chiller from plant B, Santa Catarina, Brazil.
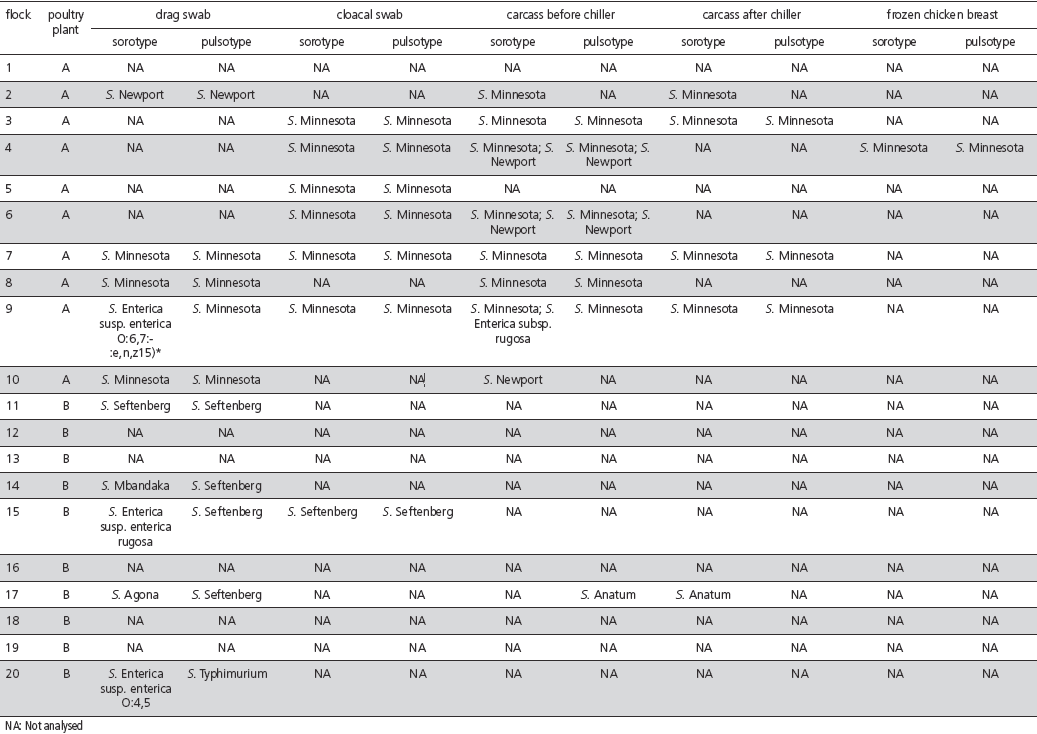
Serotyping and subtyping results of the strains isolated at processing each step are consolidated in Table 2.
Table 2 – Serotyping and PFGE pulsotypes of the strains isolated at different processing steps in two poultry processing plants located in states of Mato Grosso do Sul (plant A) and Santa Catarina (plant B), Brazil.
Ecological and regional geographic factors, such as climate, wild bird and reptile populations attracted by crops around broiler farms and processing plants, and the dominance of a specific Salmonella Minnesota cluster, present since 2010 in the evaluated region and at all poultry production steps, influenced the efficacy of the applied Salmonella prevention and control programs. This indicates that preventing the contamination of chicken meat with Salmonella poultry meat is more challenging in processing plants located in the state of Mato Grosso do Sul than in the state of Santa Catarina.
Conclusions
Considering the food safety of frozen chicken breast, the Salmonella prevention and control program adopted by the company was more effective in processing plant located in the state of Santa Catarina (plant A) than in that located in the state Mato Grosso do Sul state (plant B). This result is related with the strains prevalent in each region. In particular, the Salmonella Minnesota strain genotyped in the plant of Mato Grosso do Sul and the strains isolated from environmental samples in 2012 in the same broiler processing plant belong to a single cluster, confirming the dominance and persistence of this clone over time.
Specific tools of Salmonella prevention and control programs should be chosen and adapted to local conditions in order to reduce the food safety risks of poultry meat products throughout the production chain.
Other risk factors should be evaluated in future studies, including biofilm formation capacity of the identified Salmonella Minnesota cluster, its resistance to biocides and specific environmental conditions, its antibiotic resistance profile, the presence of pathogenicity genes, and the effective contribution of reptiles and wild birds as sources of transmission.
This article was originally published in Revista Brasileira de Ciencia Avícola. vol.19 no.1 Campinas Jan./Mar. 2017. http://dx.doi.org/10.1590/1806-9061-2016-0371. This is an Open Access article distributed under the terms of the Creative Commons Attribution License. 













.jpg&w=3840&q=75)