There is little doubt that raw milk is the most scrutinized agricultural commodity. Appendix N of the Pasteurized Milk Ordinance requires that all bulk tank milk be tested for beta-lactam antibiotics. This document also sets the regulatory levels for somatic cell counts (SCC) and bacteria levels in raw bulk tank milk. Milk processors are required to perform the regulatory testing. Many progressive milk processors provide premiums for high quality milk (low SCC and bacteria counts). From the time milk is harvested from a cow's udder, milk quality cannot improve. While SCC will not change after harvest, there are numerous reasons bacterial levels can be increased. Ideally, bacteria levels within the udder are low and additional bacterial contamination is minimized. When bacteria counts are elevated above acceptable levels, both quantitative and qualitative analyses can help pinpoint the causes.
Sources of Bacteria in Raw Milk
Milk is synthesized by cells within the mammary gland and is virtually sterile when secreted into the alveoli of the udder. Beyond this stage of milk production, bacterial contamination can generally occur from three main sources; within the udder, outside the udder, and from the surface of equipment used for milk handling and storage. Cow health, environment, milking procedures and equipment sanitation can influence the level of microbial contamination of raw milk. Equally important is the milk holding temperature and length of time milk is stored before testing and processing that allow bacterial contaminants to multiply. All these factors will influence the total bacteria count (SPC) and the types of bacteria present in raw bulk tank milk.
Microbial Contamination From Within the Udder
Raw milk as it leaves the udder of healthy cows normally contains very low numbers of microorganisms and generally will contain less than 1000 colony-forming units of total bacteria per milliliter (cfu/ml). In healthy cows, bacterial colonization within the teat cistern, teat canal, and on healthy teat skin does not significantly contribute total numbers of bacterial neither in bulk milk, nor to the potential increase in bacterial numbers during refrigerated storage. This natural flora of the cow generally will not influence the SPC, PI, LPC, or Coliform counts.
While the healthy udder should contribute very little to the total bacteria count of bulk milk, a cow with mastitis has the potential to shed large numbers of microorganisms into her milk. The influence of mastitis on the total bacteria count of bulk milk depends on type of bacteria, the stage of infection and the percent of the herd infected. Quarters from infected cows have the potential to shed in excess of 10,000,000 bacterial cfu/ml of milk produced. Mastitis organisms found to most often influence the total bulk milk bacteria counts are Streptococci (primarily Strep agalactiae and Strep uberis) although other mastitis pathogens have the potential to influence the bulk tank count as well. Staphylococcus aureus is not thought to be a frequent contributor to total bulk tank counts although counts as high as 60,000/ml have been documented.
While Staph aureus and Strep ag are rarely found outside of the mammary gland, environmental mastitis pathogens (Strep uberis and coliforms) can occur in milk as a result of other contributing factors such as dirty cows, poor equipment cleaning and/or poor cooling. Increases in SCC can sometimes serve as supportive evidence that mastitis bacteria may have caused increases in the bulk tank counts. This correlation seems to apply more for Streptococci than for Staph aureus.
Correlations between increases in somatic cell counts and other environmental mastitis organisms, including coliform bacteria, and coagulase-negative Staphylococci, were found to be poor as well. Staph aureus and Strep ag do not grow significantly on soiled milking equipment or under conditions of marginal or poor milk cooling. In general, mastitis organisms will not influence PI or LPC though in some cases of coliform mastitis, Coli counts may be elevated.
Microbial Contamination From Outside the Udder
The exterior of the cows udder and teats can contribute microorganisms that are naturally associated with the skin of the animal as well as microorganisms that are derived from the environment in which the cow is housed and milked. In general, the direct influence of natural inhabitants as contaminants in the total bulk milk count is considered to be small and most of these organisms do not grow competitively in milk. Of more importance is the contribution of microorganisms from teats soiled with manure, mud, feeds or bedding.
Teats and udders of cows inevitably become contaminated while they are lying in stalls or when allowed in dirty lots. Used organic bedding has been shown to harbor large numbers of microorganisms often exceed 100,000,000 to 10,000,000,000 per gram of bedding. Organisms associated with bedding materials that contaminate the surface of teats and udders include streptococci, staphylococci, spore-formers, coliforms and other Gram-negative bacteria. Both thermoduric and psychrotrophic strains of bacteria are commonly found on teat surfaces indicating that contamination on the outside of the udder can influence PI, LPC, and Coli counts.
The influence of dirty cows on total bacteria counts depends on the extent of soiling of the teat surface and the udder prep procedures employed. Milking heavily soiled cows could potentially result in bulk milk counts exceeding 10,000 cfu/ml. Several studies have investigated premilking udder hygiene techniques in relation to the bacteria count of milk. Generally, thorough cleaning of the teat with a sanitizing solution (predip) followed by thorough drying with a clean towel is effective in reducing the numbers of bacteria in milk contributed from soiled teats.
Microbial Contamination from Equipment Cleaning and Sanitizing Procedures
The degree of cleanliness of the milking system probably influences the total bulk milk bacteria count as much, if not more, than any other factor. Milk residue left on equipment contact surfaces supports the growth of a variety of microorganisms. Organisms considered to be natural inhabitants of the teat canal and teat skin are not thought to grow significantly on soiled milk contact surfaces or during refrigerated storage of milk. This generally holds true for organisms associated with contagious mastitis (Staph aureus and Strep ag) though it is possible that certain bacteria associated with environmental mastitis (coliforms) may be able to grow significantly. In general, bacteria from environmental contamination (bedding or manure) are more likely to grow on soiled equipment surfaces. Water used on the farm might also be a source of bacteria, especially psychrotrophs, which could seed soiled equipment.
Cleaning and sanitizing procedures can influence the degree and type of bacterial growth on milk contact surfaces by leaving behind milk residues that support growth, as well as by setting up conditions that might select for specific microbial groups. Even though equipment surfaces may be considered efficiently cleaned with hot water, more resistant bacteria (thermodurics) may endure in low numbers. If milk residue is left behind (milk stone) growth of these types of organisms, although slow, may persist. Old cracked rubber parts are also associated with higher levels of thermoduric bacteria. Significant build-up of these organisms to a point where they influence the total bulk tank count may take several days to weeks though increases would be detected in the LPC.
Less efficient cleaning, using lower temperatures and/or the absence of sanitizers tends to select for the faster growing, less resistant organisms (psychrotrophs), principally Gram-negative rods (coliforms and Pseudomonas) and some Streptococci. This will result in a high PI and in some case an elevated LPC. Effective use of chlorine or iodine sanitizers has been associated with reduced levels of psychrotrophic bacteria that cause high PI counts. Psychrotrophic bacteria tend to be present in higher bacteria count milk and are often associated with neglect of proper cleaning or sanitizing procedures and/or poorly cleaned refrigerated bulk tanks.
Milk Storage Temperature and Time
Refrigeration of raw milk, while preventing the growth of non-psychrotrophic bacteria, will select for psychrotrophic microorganisms that enter the milk from soiled cows, dirty equipment and the environment. Minimizing the level of contamination from these sources will help prevent psychrotrophs from growing to significant levels in the bulk tank during the on-farm storage period or at the processing plant. In general these organisms are not thermoduric and will not survive pasteurization. The longer raw milk is held before processing (legally up to 5 days), the greater the chance that psychrotrophs will increase in numbers. Holding milk near the PMO legal limit of 45°F allows much quicker growth than milk held below 40°F. Although milk produced under ideal conditions may have an initial psychrotrophic population of less than 10% of the total bulk tank count, psychrotrophic bacteria can become the dominant bacteria after 2 to 3 days at 40°F, resulting in a significant influence on PI counts. Colder temperatures 34-36°F will delay this shift, though not indefinitely.
Under conditions of poor cooling with temperatures greater than 45°F, bacteria other than psychrotrophs are able to grow rapidly and can become predominant in raw milk. Streptococci have historically been associated with poor cooling of milk. These bacteria will increase the acidity of milk. Certain bacteria are also responsible for a "malty defect" that is easily detected by its distinct odor. Storage temperatures greater than 60°F tend to select for these types of contaminants. The types of bacteria that grow and become significant will depend on the initial contamination of the milk.
Once milk leaves the farm, raw milk handling as well as the sub-sample collected by the milk hauler is beholden to the same sets of rules. If the raw milk or the sample used to run the regulatory tests are maintained at the proper temperature, bacterial counts can be significantly altered. Since most bulk tank milk is commingled in an over-the-road milk truck, the only way to determine each producer's contribution to the commingled milk is the sub-sample collected at pickup. Sample integrity must be maintained from the original bulk tank of milk, through hauling to the milk processor, and eventually through the end of the diagnostic procedures. As more milk processors are using bacteria counts within their premium programs, competent subsample handling is essential. In addition, much like SCC, one or two samples within a 30-day span are inadequate to provide a proper measure of a farm's management practices.
Tests For Bacteria in Raw Bulk Tank Milk
The Standard Plate Count (SPC) is used extensively in both regulatory and premium testing programs. In addition to the SPC, raw milk can be subjected to a number of other bacteriological tests that are used as indicators of how that milk was produced. These tests may be included in determining eligibility for premium payments or they may be used only as an added quality assurance tool. The bacteriological tests most often used in addition to the SPC are the Preliminary Incubation Count (PI), the Lab Pasteurization Count (LPC) and the Coliform Count. While the SPC gives an estimated count of the total bacteria in a sample the PI, LPC and Coli Count select for specific groups of bacteria that are associated with poor milking practices. Results of these testing procedures are used to help identify potential problems that may not be detected by the SPC.
Standard Plate Count
The Standard Plate Count (SPC) of raw milk gives an indication of the total number of aerobic bacteria present in the milk at the time of pickup. Milk samples are plated in a semi-solid nutrient media and then incubated for 48 hrs at 32°C (90°F) to encourage bacterial growth. Single bacteria (or clusters) grow to become visible colonies that are then counted. All plate counts are expressed as the number of colony forming units per milliliter (cfu/ml) of milk. Newer filmsbased tests have allowed for automation of this procedure. The nutrient agar system remains the gold standard.
Aseptically collected milk from clean, healthy cows generally has an SPC less than 1,000 cfu/ml. Higher counts suggest bacteria are entering the milk from a variety of possible sources. Though it is impossible to eliminate all sources of contamination, counts of less than 5,000 cfu/ml are possible, while counts of 10,000 cfu/ml should be achievable by most farms. The most frequent cause of a high SPC is poor cleaning of the milking system. Milk residues on equipment surfaces provide nutrients for growth and multiplication of bacteria that contaminate the milk at subsequent milk times. Other procedures that can elevate bulk-tank SPC are milking dirty udders, maintaining an unclean milking and housing environment, and failing to rapidly cool milk to less than 40°F. On rare occasions, cows with mastitis can shed infectious bacteria into the milk can also cause a high SPC. In these circumstances, a concurrent elevation in SCC should be evident.
Preliminary Incubation Count
The Preliminary Incubation Count (PI) reflects milk production practices. This procedure involves holding the milk at 55°F for 18 hours prior to plating. This step encourages the growth of groups of bacteria that grow well at cool temperatures (psychrotrophs). Bacteria in the incubated sample are counted with the SPC procedure and compared to the SPC from the same sample to determine if a significant increase has occurred. PI counts are generally higher than the SPC. Counts with a 3-4-fold increase are considered significant. Some consider counts greater than 50,000 cfu/ml to be of concern regardless of the SPC, though in some cases the counts may be equal and in rare cases the PI may be lower.
High PI counts are most often associated with a failure to thoroughly clean and sanitize either the milking system or the cows. Bacteria considered to be natural flora of the cow, including those that cause mastitis, are not thought to grow significantly at the PI temperature. However, PI equal to or slightly higher than a high SPC (greater than 50,000 cfu/ml) may suggest that the high SPC is possibly due to mastitis pathogens. Marginal cooling (milk held at temperatures over 40°F) or prolonged storage times may also result in unacceptable PI levels by allowing organisms that grow at refrigeration temperatures to multiply.
Lab Pasteurized Count
Though most bacteria are destroyed by pasteurization, there are certain types and certain bacterial stages that are not. The Lab Pasteurized Count (LPC) estimates the number of bacteria that can survive the pasteurization process. Milk samples are heated to simulate batch pasteurization at 145°F for 30 minutes. Bacteria that survive pasteurization (thermodurics) are then counted using the SPC procedure. LPC are generally much lower than SPC, with counts greater than 300 considered high. Bacteria found in the natural flora of the cow, as well as those associated with mastitis, are generally not thermoduric. High LPC values are most often associated with a chronic or persistent cleaning failure in some area of the system or significant levels of contamination from soiled cows. Other common causes of high LPC are leaky pumps, old pipeline gaskets, inflations and other rubber parts, and milk stone deposits.
Coliform Count
The Coliform Count (Coli Count) procedure selects for bacteria that are most commonly associated with manure or environmental contamination. Milk samples are plated on a selective nutrient media that encourages the growth of coliform bacteria, while preventing the growth of others. Although coliforms are often used as indicators of fecal contamination, there are strains that commonly exist in the environment. Coliforms may enter the milk supply as a consequence of milking dirty cows or the claw becoming soiled with manure during milking. Generally, counts greater than 100 cfu/ml would indicate poor milking hygiene or other sources of contamination. Higher coliform counts more often result from dirty equipment and in rare cases result from milking cows with environmental coliform mastitis.
Summary of Test Results and Sources of Bacterial Contamination in Raw Milk
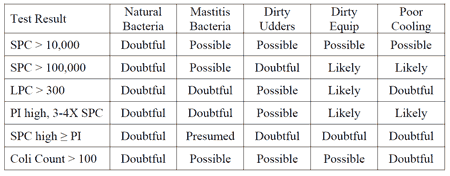 Take Home Messages
Take Home Messages
- Bacterial contamination of raw milk can generally occur from three main sources; within the udder, outside the udder, and from the surface of equipment used for milk handling and storage.
- The bacteriological tests used most often are the Standard Plate Count (SPC), the Preliminary Incubation Count (PI), the Lab Pasteurization Count (LPC) and the Coliform Count.











