1. Introduction
In the US, 30 serovars of Salmonella enterica causing foodborne illness persist at an incidence in the general food supply to warrant continuous survey [1]. Several serovars are of interest for their association with eggs and poultry products. Phenotype microarray (PM) data were accumulated for the top 4 serovars of Salmonella enterica linked to chicken (Typhimurium, Enteritidis, Heidelberg and Infantis) and one associated with turkey (Senftenberg) [1,4]. During accessioning of PM datasets, we observed that respiratory activity was present or absent in PM10 A03 (pH 4.5). The disparities between strains appeared large in comparison to other wells in the pH gradient (PM10 A01 e PM10 A12). The acid tolerance response is generally accepted as a fundamental characteristic of the Salmonellae that facilitates survival of the pathogen in a range of acidic environments, both internal and external to avian and mammalian hosts [9]. The acid tolerance response is complex in its induction, has variable determinants and has been associated with multiple inducers, genes and regulatory pathways [3,14,20,21]. Within the food industry, survival of Salmonella in acidic conditions is of great concern because low pH is used as a preservative [10,15,18]. We thus wanted to know if isolates that varied in acid tolerance had other linked metabolic differences. Also of value was the development of a general statistical strategy for processing large amounts of information derived from PM analysis of Salmonella enterica wildtype isolates.
2. Materials and methods
2.1. Description of bacteria
Bacteria used for this study were obtained from a variety of proprietary sources and chosen for inclusion based on a known association with poultry over a period of 3 years. S. enterica serotypes represented are Typhimurium (ST), Enteritidis (SE), Heidelberg (SH), Infantis (SI) and Seftenberg (SS). The first 4 serotypes have a strong association of foodborne illness and chicken products, whereas the last one is more associated with turkey [1]. Each serovar had multiple independent isolates, noted by a number following the serotype designation. Thus, SE7 was the 7th independent isolate of serovar Enteritidis assayed. Some isolates were repeatedly assayed to determine variation occurring between experiments over time. For example, SE7 was used as a primary control strain and it was assayed 10 times, and thus datasets were numbered SE7-1 through SE7-10 in tables. Other isolates that were assayed more than once include SE5 (4 runs), SI4 (2 runs), SI6 (2 runs) and SH11 (2 runs). Isolates were stored at 80 °C in 15% glycerol pending analysis.
2.2. Parameters of analysis
Runs were conducted in Omnilog SN248 (Biolog, Inc., Hayward, CA, USA) at 42 °C, which is the normal body temperature of poultry. Data were collected over 48 h. Isolates were revived from frozen stock onto Biolog universal growth (BUG) agar prepared with 5% sheep blood according to the provider's directions (Biolog, Inc.). Plates were incubated for 16 h at 37 °C and then placed at 4 °C for at least 1 h. Each isolate was processed individually in order to keep the temperature of media near 4 °C, which aids in keeping background respiratory activity minimized. Cells were suspended in IF-0 without dye (Biolog, Inc.) to a 42% suspension and then transferred according to PM directions to IF-0 with dye (PM1 through 8) or IF-10 with dye (PM 9 and 10). Suspensions were returned to 4 °C. One suspension at a time was plated to PM1-10, sealed with plastic and then kept at 4 °C until information had been entered and the machine was ready to load and run. Respiratory activity results in an irreversible dye reaction that is digitally captured every 15 min and converted to respiratory units (RU) by proprietary software (Biolog, Inc.). Average height was used for analysis of datasets
2.3. Statistical analyses
2.3.1. Determination of background respiratory units (RU)
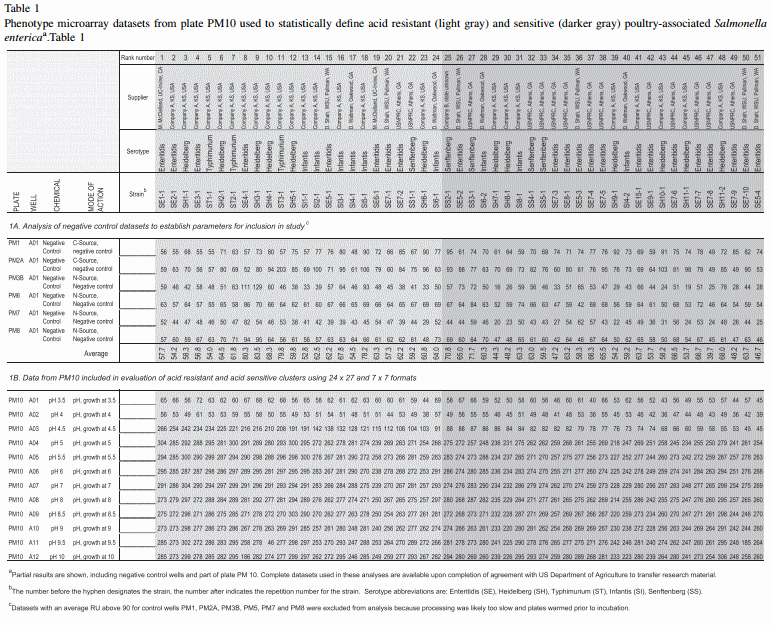
The A01 negative control wells from plates PM1, PM2A, PM3B, PM6, PM7 and PM8 were used to calculate average background RU from an initial 57 datasets (Table 1A). Any individual dataset that had an average for its 6 negative control wells exceeding background RU was excluded from analysis. Background RU is very sensitive to technical aspects of loading and handling plates and methods used to inhibit background RU as described above must be followed to maintain quality of any one run.
2.3.2. Initial cluster analysis and calculations for assessing significance
To begin analysis, data were first partitioned into two clusters based on results from 51 qualifying datasets as ranked from greatest to least by RU at PM10 A03, pH 4.5. This initial cluster is referred to as the 24 x 27 format, because 24 strains appeared acid-resistant and 27 strains appeared sensitive (Table 1B). Three types of calculations were performed on the 24 x 27 formatted clusters, and these were filtered in order to observe the most likely differences in phenotype (Table 2). First, standard deviation (stdev) was calculated using the Excel STDEV.S function for each of the 950 chemicals. Chemicals that did not have at least one stdev of at least 24 RU across all data points were essentially homogeneous and of no significance. Then, the probability value ( p) by Student's t-Test was calculated using Excel. Any chemical that did not have a p value less than 0.030 was excluded, or filtered out, as being significantly different. For the 24 x 27 format, tails ¼ 1, type ¼ 2 and for the paired 7 x 7 format, tails ¼ 1, type ¼ 1.
Effect size (ES) was also determined by calculating the average of values for two clusters and then dividing the difference of the averages by the standard deviation [5]. ES conveys the magnitude of difference between clusters, and the greater the ES, the greater the magnitude of the difference. ES assesses the magnitude of an effect based on absolute values, and guidelines suggest that minor effects are in the range of ±0.2 to 0.4, moderate effects are in the range of ±0.4 to 0.8, and major effects exceed ±0.8 [5,6]. For these analyses, a positive ES was associated with acid resistance and a negative ES was associated with acid sensitivity, because the formula used for each chemical was AR (average of the acid-resistant cluster) e AS (average of the acid-sensitive cluster) divided by the stdev calculated from AR and AS values combined. Only absolute ES values of ≤ –0.50 or ≥0.50 were considered significant.
2.3.3. Additional cluster analysis for minimizing false negatives
A second clustering strategy was used and it also filtered datasets in order of stdev, p value and ES as done for the 24 x 27 format (Table 2). Hierarchical, or subset, clustering included 7 each of the most extreme datasets, namely the top half and bottom half of the first and fourth quartiles. This grouping is referred to as the 7 x 7 format. Hierarchical clustering is more likely to increase false-positive results, but is less likely to miss a phenotype of interest [5].
3. Results
3.1. Initial assessment of statistically significant differences between putatively acid-resistant and acid-sensitive clusters
The average of all 6 control wells from 57 samples was 65.5 RU and the standard deviation was 24.61 RU (Table 1A). Thus, any dataset with an average RU of the average plus one stdev, namely 90 RU or above, fell outside of the range of acceptable background, and 6 datasets were removed from the study. The 51 datasets that had acceptable background RU of no more than 90 RU were then divided into two clusters, with 24 acid-resistant isolates having an average RU at pH 4.5 above 90 RU and 27 below (Table 1A). Application of Student's t-test to all data within each cluster gave a p value (tails 1, type 2) of 0.001 when the clusters were compared.
Effect size (ES) was also calculated to assess the magnitude of the difference associated with a very low p value. Results indicated that the overall difference between the two groups of all inclusive datasets was negligible because ES was 0.028. These results suggest that meaningful differences might exist within the entire dataset, but only a minority of chemicals were likely to be significantly different.
3.2. Parameters further defining significant differences between acid-resistant and acid-sensitive clusters
Based on initial assessments, the final parameters for defining differences between acid-resistant and -sensitive clusters as significant required: i) an stdev ≥24 RU (one stdev), ii) p ≤ 0.030, ii) and iii) a moderate ES with absolute value ≥0.5. Gradients in PM9 and 10, other than PM10 A03 used to define clusters, were evaluated somewhat differently because two concentrations in a row had to meet the definition of significance to be considered especially robust. The extra parameter corrects for minor pipetting differences between runs. Given the inherent variability observed between repetitions, e.g. variation in values for 10 repetitions of SE7 (Tables 1A and 1B), findings were described as significant only after applying the different layers of analyses.
3.3. Analysis of variability between runs
3.3.1. Cluster analysis of 24 putatively acid-resistant datasets compared to 27 acid-sensitive (24 x 27 format)
Data in Table 1 were ranked in order as determined by values at pH 4.5 (well PM10 A03), signifying greatest to least acid resistance. Ranking for the 10 repetitions of SE7 placed the strain from 20th to 50th, 4 repetitions of SE5 placed from 15th to 51st, 2 of SI4 placed it at 17th and 40th and SI6 placed 24th and 28th. Thus, 10 different runs of a single isolate of Enteritidis (SE7) clustered eight times as acid-sensitive and two times as resistant, and other serovars had a similar spread. These results suggest that stringent analysis is required to detect stable phenotypic differences and that some variation is likely to occur between colonies and runs. Cluster analysis for assigning acid resistance within the 24 x 27 format might be more robust if the observed biological parameters for SE7 were used to set background RU in future studies. For example, an RU of at least 115 rather than 90 may be the better parameter for setting a boundary
3.3.2. Increasing stringency of comparison by hierarchical cluster analysis of the 7 most and least acid-resistant groups (7 x 7 format)
Comparing groups of the 7 most and least acid-resistant datasets examines the top ½ of the two most extreme quartiles. By comparing datasets from both extremes, a broader overview of potentially significant results was expected, because false-positives would increase but false-negatives would decrease. Results indicated that the 7 x 7 format found many more differences between the selected subsets (Table 2). The 24 x 27 format showed that 75 individual wells of 950 test wells appeared significantly different, while the 7 x 7 format showed 220 wells. Thus, between 7.89% and 23.16% of metabolites showed some association with either acid resistance or sensitivity.
3.3.3. Acid resistance was not linked to virulence
SE8 is a sefD mutant of parent SE7, and it placed 35th. Previous complementation of SE8 with a high copy plasmid to produce SE9 enabled constitutive expression of sefD, altered cell surface properties and resulted in attenuation, but placement only changed to 42nd in rank. The sefD mutant of SE7 was exceptionally virulent in previous studies and it would kill healthy hens at an incidence higher than the wild-type parent; however, the complemented mutant was greatly attenuated [22]. Thus, acid resistance was not linked to virulence attributes of serovar Enteritidis in this study. It is possible that other types of mutants impact regulatory circuits that do link acid resistance to virulence.
3.4. Results from cluster analysis of datasets by class of chemical
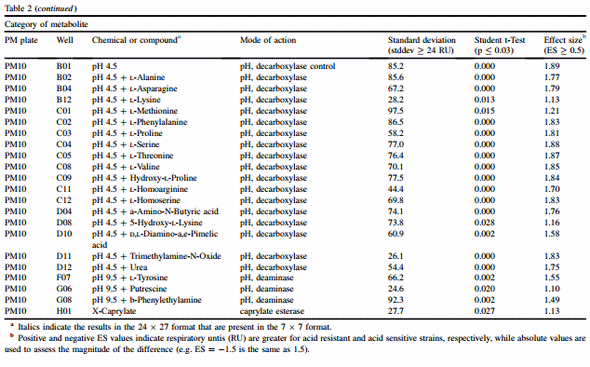
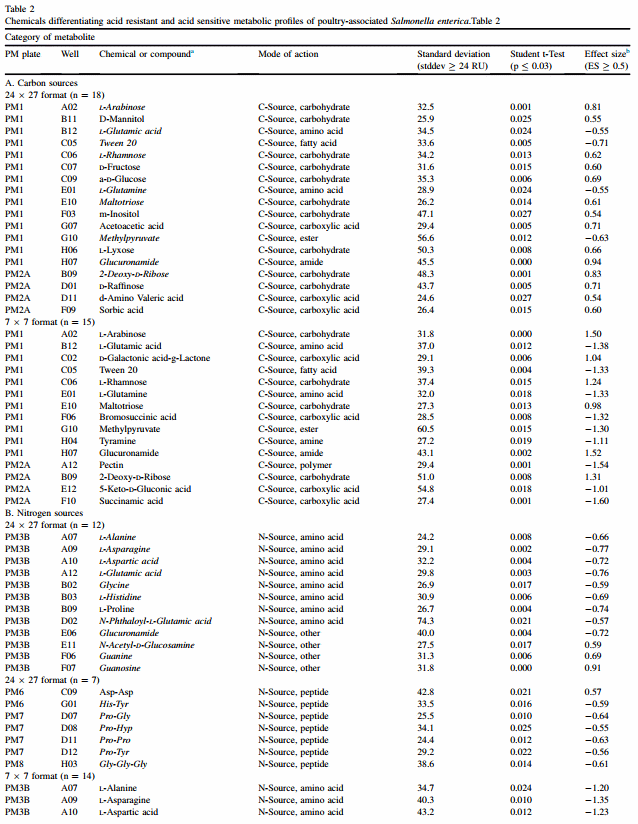
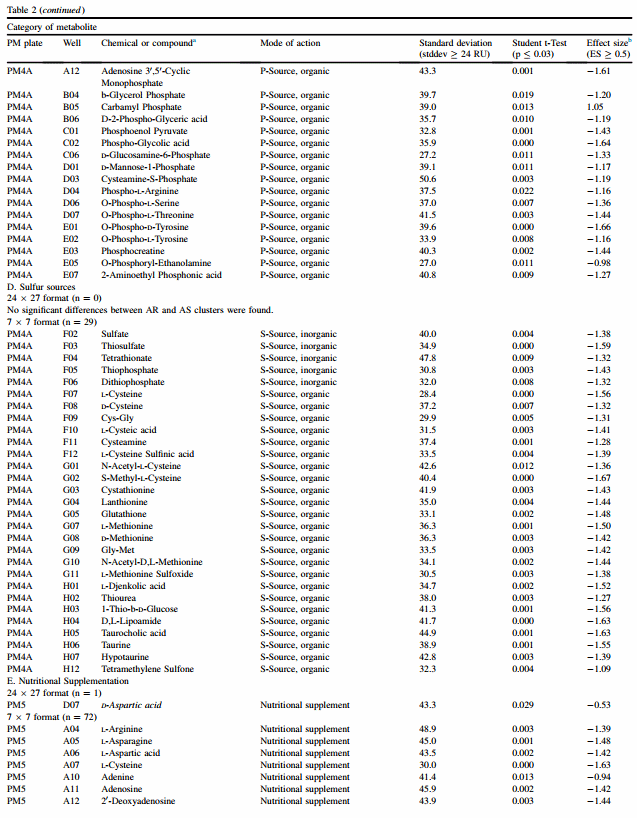
For both 24 x 27 and 7 x 7 formats, results are discussed according to class of chemical for plates PM1 e PM8. Classes were C-source (PM1 and PM2), N-source (PM3, PM6, PM7 and PM8), P-source (PM4), S-source (PM4) and nutritional supplementation (PM5). PM9 and PM10 are composed mostly of different types of gradients and each one is discussed individually. Table 2 summarizes findings for both formats, and chemicals found in common between the two formats are italicized.
3.4.1. C-source (PM1 and PM2)
Analysis using a 24 x 27 format found significant differences between acid resistance and sensitive clusters for 18 chemicals out of 190, whereas the 7 x 7 format found 15. Three (3) chemicals were in the top ten by ES for both formats, and these were Tween 20, L-arabinose and 2-deoxy-Dribose acid (Table 2).
3.4.2. N-source (PM3, PM6, PM7 and PM8)
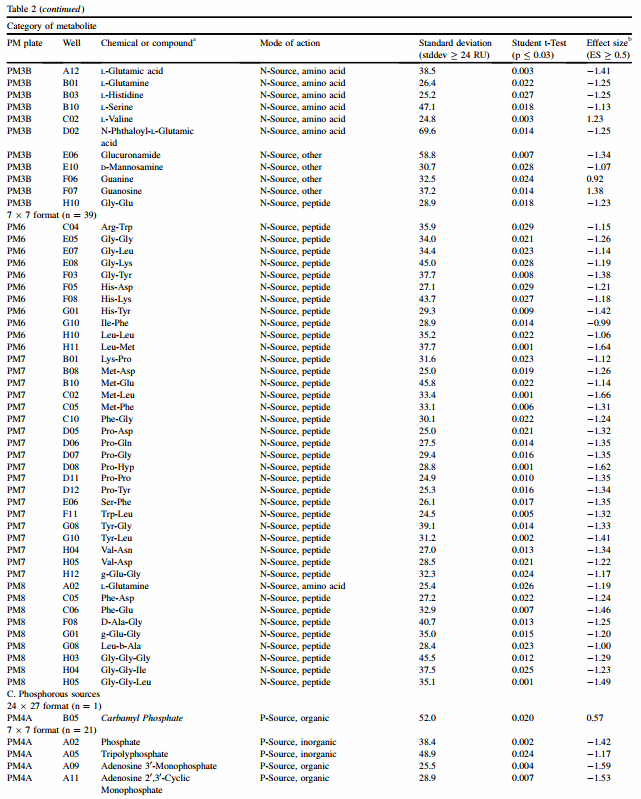
In Table 2, N-source chemicals in PM3B are separated from peptides tested in PM6, 7 and 8. The 24 x 27 format found a total of 19 N-source chemicals out of 381 that appeared to be metabolized differently by acid-resistant and -sensitive clusters. For the 24 x 27 format, guanosine was the chemical with the greatest ES value for acid-resistant strains (ES ¼ 0.91), and this value was even greater when analyzed in the 7 x 7 format (ES ¼ 1.38). Acid-sensitive strains appeared to metabolize L-asparagine better than resistant ones (ES ¼ 0.77), and the order of magnitude in the 7 x 7 format was again greater (ES ¼ 1.35).
Application of the 7 x 7 format detected 53 chemicals that differed between the two clusters (Table 2). For the 7 x 7 format, 3 diamino acids, namely Met-Leu, Leu-Met and ProHyp, were the 3 chemicals with the highest absolute ES values for acid-sensitive strains. For the 53 N-source chemicals identified as having significant differences between clusters, 91 (97.8%) were associated with acid sensitivity and only 2 with acid resistance (data not shown). Guanosine and LvValine were the only two chemicals associated with significantly greater metabolism by resistant strains (ES ¼ 1.38 and 1.23, respectively). This suggests that chemicals metabolized by acid-resistant strains may be a smaller set than those used by acid-sensitive strains.
3.4.3. P-source (PM4)
The 24 x 27 format found one significant difference between clusters, while the 7 x 7 format found 21 (Table 2). Carbamyl phosphate was found by both formats with a higher RU for the acid-resistant cluster, but its effect was moderate (ES ¼ 0.57 and 1.05 respective to 24 x 27 and 7 x 7 formats). As with results from nitrogen sources, these results suggest that acid-sensitive strains metabolize a greater range of compounds than do acid-resistant strains.
3.4.4. S-source (PM4)
No sulfur sources were found by the 24 x 27, with a significant difference between acid-resistant and -sensitive clusters. The 7 x 7 format found that 29 of 35 sulfur sources met criteria for supporting metabolism of acid-sensitive strains (Table 2). Thus, it appears that acid-sensitive strains metabolize a wide range of sulfur sources. D,L-ethionine was the only sulfur source that appeared neutral in its impact for acid-resistant and -sensitive strains (PM4A G06).
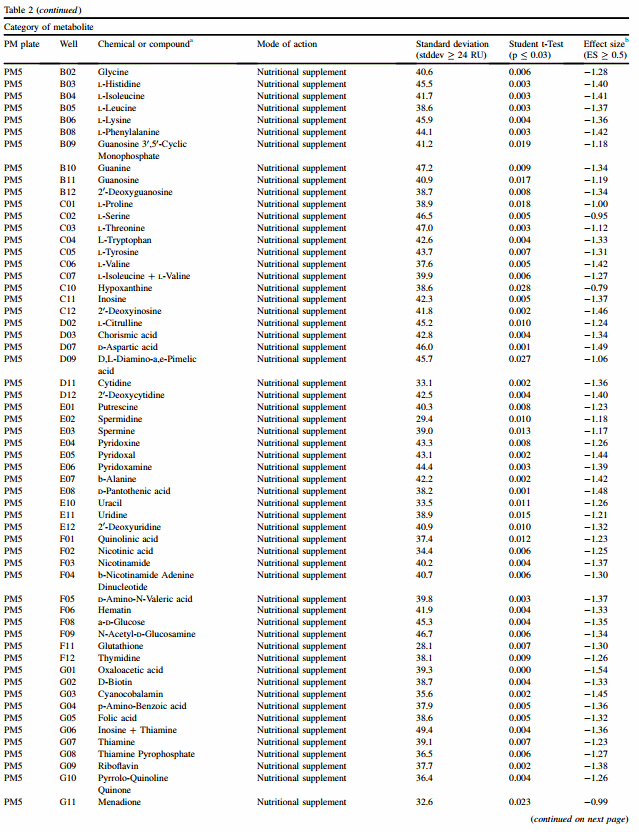
3.4.5. Nutritional supplementation (PM5)
Nutritional supplementation with 94 different chemicals did not appear to significantly alter RU between clusters using the 24 x 27 format, with the exception of marginal results for Daspartic acid (PM5 D07, ES ¼ 0.53) (Table 2). In contrast, the 7 x 7 format found 72 significant differences, and all were associated with higher RU of datasets in the acid-sensitive cluster.
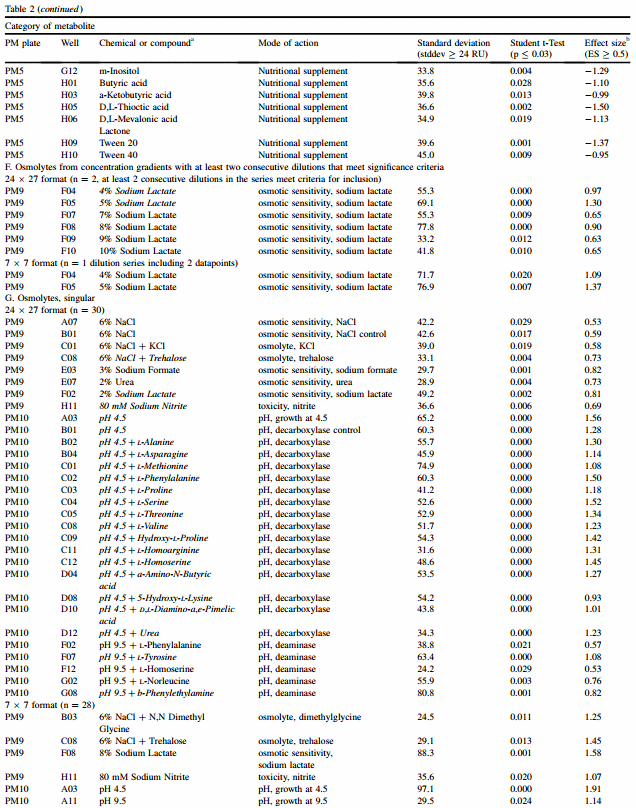
3.4.6. Gradients of osmolytes (PM9 and PM10)
Fourteen different gradients are included on PM9 and PM10, and the pH gradient as shown in Table 1B is also included. Tolerance to different concentrations of sodium lactate was significantly better for the acid-resistant cluster, as detected within the 24 x 27 format. The 7 x 7 format also detected sodium lactate resistance for the cluster of isolates that were acid-resistant.
3.4.7. Singular osmolytes (PM9 and PM10)
In addition to the pH gradient, PM10 has 35 chemicals prepared at pH 4.5 (B02 e D12), with PM10 B01 serving as a decarboxylase control. Respectively, 15 and 17 chemicals tested at pH 4.5 were found by the 24 x 27 and 7 x 7 formats to be associated with significantly higher RU by the acid-resistant cluster (Table 2). While most chemicals were shared by both formats, the 7 x 7 format included L-lysine and trimethylamine-N-oxide. Respective to the two formats, 5 and 3 chemicals at pH 9.5 were found to differ between the two clusters, and again acid resistance was the phenotype that had higher RU values (data not shown). Individual growth conditions for 6% sodium chloride, 8% sodium formate, urea, sodium lactate and 80 mM sodium nitrite were also found to be significant and again favored acid resistance. These results suggest that the acid-resistant isolates may have some resistance to a range of other conditions, but that acid resistance at pH 4.5 was predominant, as measured by ES.
4. Discussion
We applied a layered statistical analysis using two types of cluster analysis combined with standard deviation, p value and effect size to assess metabolic traits linked to the acid tolerance response of S. enterica. Type II error, which is the failure to detect a difference that is present, is more likely within the 24 x 27 format, and false-negative error is in large part due to obfuscation arising from significant inherent variation occurring between sample runs. The 7 x 7 format increased the possibility of making a Type I false-positive error, which claims a difference to be present when it is not. Part of the analysis was to set limits for what constituted acceptable background, and thus 6 datasets were excluded. The spurious presence of background RU explains in part why proof of an effect requires biological experimentation for each chemical of interest. The approach used here is suggested as being most appropriate for analysis of field isolates. Repetition across time is advised for analyzing complex phenotypic traits of field isolates within any microarray format [7,16,23]. More than 3 repetitions may not necessarily be needed for mutational studies, because genetic engineering aids in fixing and exaggerating phenotypic differences, so that differences are easier to discern [19]. Nor is similar repetition needed to observe very clear absence or presence of discrete phenotypes, such as the ability to metabolize D-serine, which was first seen in a naturally occurring mutant of S. enterica by phenotype microarray and later traced to the presence of a single nucleotide difference [11]. As referenced in the Introduction, the nature of the phenotype under investigation, namely acid resistance or sensitivity, is very complex and subject to subtle regulatory clues; thus, exceptionally stringent analysis was required to observe differences. An area of future research is to design media to differentiate acid-resistant and -sensitive populations for routine surveillance. For example, a minimal media for acid-resistant Salmonella might include L-arabinose (C-source), guanosine (N-source), carbamyl phosphate (P-source) and D,L-ethionine (S-source). A minimal media for sensitive populations might contain L-glutamic acid (C-source), L-asparagine (N-source), phosphate (P-source) and tetrathionate (S-source). Methods used to isolate Salmonella already include the use of tetrathionate media as well as other media made from tryptic digests of casein [2]. Thus, another area of research is to determine if the acid-resistant and -sensitive populations listed differ in recovery from these media.
Acid resistance was strongly associated with tolerance at pH 4.5, and the 24 x 27 format detected 15 of the 17 chemicals detected by the 7 x 7 format for PM10 tests conducted at pH 4.5. Unexpected findings were that resistance to sodium lactate might be linked to acid-resistant S. enterica. Crossover effects have been observed by others under experimental settings [10]. Sodium lactate and acidic conditions are commonly used as food preservatives. Thus, it may be as important to understand risk factors associated with resistance developing to multiple antimicrobial compounds used in food preparation as it is to investigate evolution of multiply antibiotic-resistant organisms causing clinical illness. Another area of concern is that resistance to urea, sodium chloride, sodium nitrite and pH 9.5 might be emerging, as suggested by results appearing significant for singular osmolytes.
The chemicals listed in Table 2 as being differentially metabolized by populations that differ in acid resistance have some support from former findings. For example, L-arabinose was associated with acid resistance in this study, and this chemical may have a role in environmental signaling and suppression of Salmonella pathogenicity island 1 (SPI-1) genes [17]. Differential carbon utilization was linked to planktonic versus biofilm cells; specifically, pectin (C-source, acid-sensitive) induced biofilm formation in a human carrier isolate of serovar Typhi [13]. Another study suggested differences in metabolism between Salmonella serovars Derby and Mbandaka that included some of the chemicals observed here [12]. S. enterica can use 2-deoxy-D-ribose as a sole C-source, but Escherichia coli cannot [8], and perhaps acidresistant strains metabolize this chemical better. Further research is required to prove that the chemicals listed in Table 2 are differentially metabolized by S. enterica that varies in acid resistance, and that differences matter within the larger context of assuring the safety of food.
Conflicts of interest
None.
Acknowledgments
This research was primarily supported by ARS Project Number 6040-32000-007-D and supplementary support from ARS Project Number 6040-32420-001-00. CEVA and Aviagen® provided some travel support and collaboration under material transfer agreements, which protected rights to authorship by the USDA. Bwalya Lungu was an employee of Aviagen while participating in this research. Jean Guard, with assistance from laboratory technician Tod Stewart, conducted phenotype microarray. Collaborators contributed bacteria and essential support exceeding 3 years. We also want to thank Michael McClelland and colleagues for their contributions of isolates and continued support on genomic analyses indirectly related to this manuscript. The complete dataset for PM plates 1e10 is available upon request in spreadsheet form according to USDA guidelines for a material transfer research agreement (MTRA).
This article was originally published in Research in Microbiology 167 (2016) 745e756. http://dx.doi.org/10.1016/j.resmic.2016.06.006. This is an Open Access article under the CC BY-NC-ND license (http://creativecommons.org/licenses/by-nc-nd/4.0/).
Reference
[1] (CDC) CfDCaP. An Atlas of Salmonella in the United States, 1968e2011: laboratory-based enteric disease surveillance. In: Department of Health and Human Services (HHS). Atlanta, GA: CDC; 2013.
[2] Andrews WH, Jacobson A, Hammack T. Salmonella. Bacteriological analytical manual, food and drug administration (FDA). Gaithersburg, MD: Department of Health and Human Services (HHS); 2014.
[3] Bang IS, Kim BH, Foster JW, Park YK. OmpR regulates the stationaryphase acid tolerance response of Salmonella enterica serovar typhimurium. J Bacteriol 2000;182:2245e52.
[4] Bochner BR. Global phenotypic characterization of bacteria. FEMS Microbiol Rev 2009;33:191e205.
[5] Borenstein M. Hypothesis testing and effect size estimation in clinical trials. Annals of allergy, asthma & immunology. Off Publ ACAAI 1997; 78:5e11. quiz 2e6.
[6] Brady SM, Burow M, Busch W, Carlborg O, Denby KJ, Glazebrook J, et al. Reassess the t test: interact with all your data via ANOVA. Plant Cell 2015;27:2088e94.
[7] Chen JJ, Hsueh HM, Delongchamp RR, Lin CJ, Tsai CA. Reproducibility of microarray data: a further analysis of microarray quality control (MAQC) data. BMC Bioinform 2007;8:412.
[8] Christensen M, Borza T, Dandanell G, Gilles AM, Barzu O, Kelln RA, et al. Regulation of expression of the 2-deoxy-D-ribose utilization regulon, deoQKPX, from Salmonella enterica serovar typhimurium. J Bacteriol 2003;185:6042e50.
[9] Foster JW. Low pH adaptation and the acid tolerance response of Salmonella typhimurium. Crit Rev Microbiol 1995;21:215e37.
[10] Greenacre EJ, Brocklehurst TF. The acetic acid tolerance response induces cross-protection to salt stress in Salmonella typhimurium. Int J Food Microbiol 2006;112:62e5.
[11] Guard J, Morales CA, Fedorka-Cray P, Gast RK. Single nucleotide polymorphisms that differentiate two subpopulations of Salmonella enteritidis within phage type. BMC Res Notes 2011;4:369.
[12] Hayward MR, AbuOun M, Woodward MJ, Jansen VA. Temperature and oxygen-dependent metabolite utilization by Salmonella enterica serovars Derby and Mbandaka. PloS One 2015;10:e0120450.
[13] Kalai Chelvam K, Yap KP, Chai LC, Thong KL. Variable responses to carbon utilization between planktonic and biofilm cells of a human carrier strain of Salmonella enterica serovar Typhi. PloS One 2015;10:e0126207.
[14] Lee IS, Slonczewski JL, Foster JW. A low-pH-inducible, stationary-phase acid tolerance response in Salmonella typhimurium. J Bacteriol 1994;176:1422e6.
[15] Lianou A, Koutsoumanis KP. Evaluation of the strain variability of Salmonella enterica acid and heat resistance. Food Microbiol 2013;34: 259e67.
[16] Liu F, Kuo WP, Jenssen TK, Hovig E. Performance comparison of multiple microarray platforms for gene expression profiling. Methods Mol Biol 2012;802:141e55.
[17] Lopez-Garrido J, Puerta-Fernandez E, Cota I, Casadesus J. Virulence gene regulation by L-arabinose in Salmonella enterica. Genetics 2015; 200:807e19.
[18] Mokgatla RM, Brozel VS, Gouws PA. Isolation of Salmonella resistant to hypochlorous acid from a poultry abattoir. Lett Appl Microbiol 1998; 27:379e82.
[19] Morales CA, Guard J, Sanchez-Ingunza R, Shah DH, Harrison M. Virulence and metabolic characteristics of Salmonella enterica serovar enteritidis strains with different sefD variants in hens. Appl Environ Microbiol 2012;78:6405e12.
[20] Ryan D, Pati NB, Ojha UK, Padhi C, Ray S, Jaiswal S, et al. Global transcriptome and mutagenic analyses of the acid tolerance response of Salmonella enterica serovar typhimurium. Appl Environ Microbiol 2015; 81:8054e65.
[21] Rychlik I, Barrow PA. Salmonella stress management and its relevance to behaviour during intestinal colonisation and infection. FEMS Microbiol Rev 2005;29:1021e40.
[22] Sanchez-Ingunza R, Guard J, Morales CA, Icard AH. Reduction of Salmonella Enteritidis in the spleens of hens by bacterins that vary in fimbrial protein SefD. Foodborne Pathog Dis 2015;12:836e43.
[23] Shi L, Jones WD, Jensen RV, Harris SC, Perkins RG, Goodsaid FM, et al. The balance of reproducibility, sensitivity, and specificity of lists of differentially expressed genes in microarray studies. BMC Bioinform 2008;9(Suppl 9):S10.