Environmental distribution and genetic diversity of vegetative compatibility groups determine biocontrol strategies to mitigate aflatoxin contamination of maize by Aspergillus flavus
Author details
1 Plant Pathology Unit, International Institute of Tropical Agriculture, PMB 5320 Ibadan, Nigeria; 2 Institute for Plant Diseases, Phytopathology and Nematology in Soil Ecosystems, University of Bonn, Bonn, Germany; 3 Department of Plant Pathology, North Carolina State University, Raleigh, North Carolina, USA; 4 Department of Crop Protection and Environmental Biology, University of Ibadan, Ibadan, Nigeria; 5 USDA-ARS, Division of Plant Pathology and Microbiology, Department of Plant Sciences, University of Arizona, Tucson, Arizona, USA.
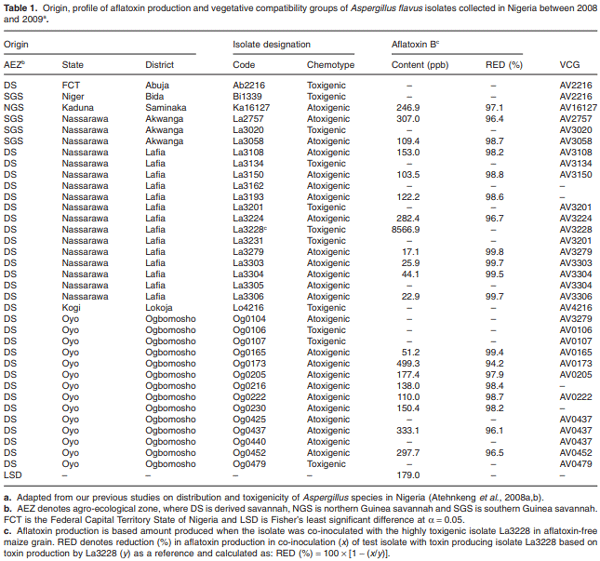
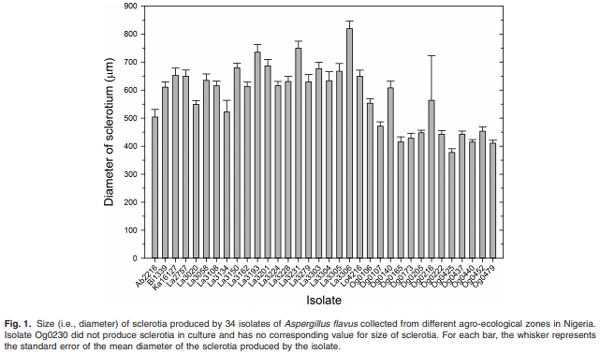
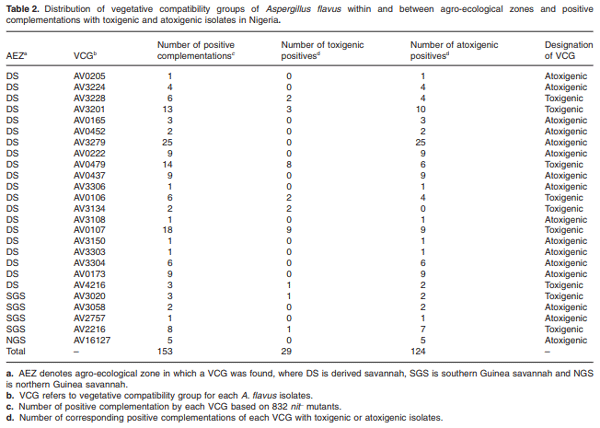
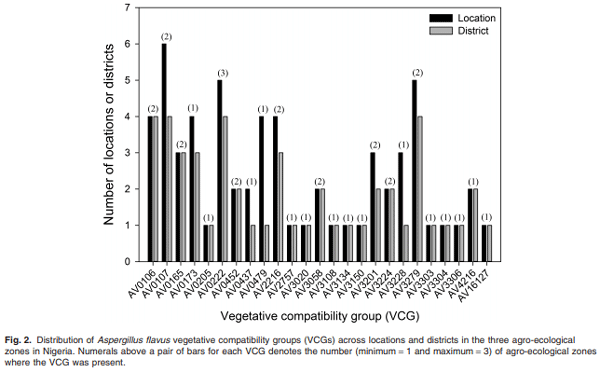
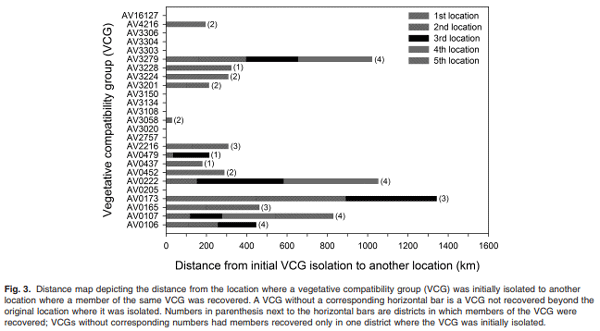
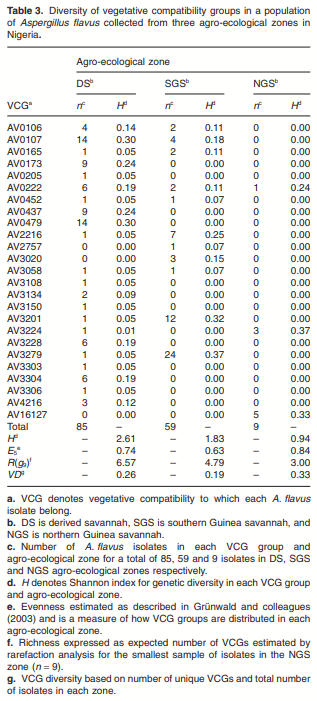
Maize infected by aflatoxin-producing Aspergillus flavus may become contaminated with aflatoxins, and as a result, threaten human health, food security and farmers’ income in developing countries where maize is a staple. Environmental distribution and genetic diversity of A. flavus can influence the effectiveness of atoxigenic isolates in mitigating aflatoxin contamination. However, such information has not been used to facilitate selection and deployment of atoxigenic isolates. A total of 35 isolates of A. flavus isolated from maize samples collected from three agroecological zones of Nigeria were used in this study. Ecophysiological characteristics, distribution and genetic diversity of the isolates were determined to identify vegetative compatibility groups (VCGs). The generated data were used to inform selection and deployment of native atoxigenic isolates to mitigate aflatoxin contamination in maize. In co-inoculation with toxigenic isolates, atoxigenic isolates reduced aflatoxin contamination in grain by > 96%. A total of 25 VCGs were inferred from the collected isolates based on complementation tests involving nitrate non-utilizing (nit− ) mutants. To determine genetic diversity and distribution of VCGs across agroecological zones, 832 nit− mutants from 52 locations in 11 administrative districts were paired with one self-complementary nitrate auxotroph tester-pair for each VCG. Atoxigenic VCGs accounted for 81.1% of the 153 positive complementations recorded. Genetic diversity of VCGs was highest in the derived savannah agro-ecological zone (H = 2.61) compared with the southern Guinea savannah (H = 1.90) and northern Guinea savannah (H = 0.94) zones. Genetic richness (H = 2.60) and evenness (E5 = 0.96) of VCGs were high across all agro-ecological zones. Ten VCGs (40%) had members restricted to the original location of isolation, whereas 15 VCGs (60%) had members located between the original source of isolation and a distance > 400 km away. The present study identified widely distributed VCGs in Nigeria such as AV0222, AV3279, AV3304 and AV16127, whose atoxigenic members can be deployed for a region-wide biocontrol of toxigenic isolates to reduce aflatoxin contamination in maize.
Atehnkeng, J., Ojiambo, P.S., Ikotun, T., Sikora, R.A., Cotty, P.J., and Bandyopadhyay, R. (2008a) Evaluation of atoxigenic strains of Aspergillus flavus as potential biocontrol agents for aflatoxin in maize. Food Addit Contam 25: 1264–1271.
Atehnkeng, J., Ojiambo, P.S., Donner, M., Ikotun, T., Sikora, R.A., Cotty, P.J., and Bandyopadhyay, R. (2008b) Distribution and toxigenicity of Aspergillus species isolated from maize kernels from three agro-ecological zones in Nigeria. Int J Food Microbiol 122: 74–84.
Atehnkeng, J., Ojiambo, P.S., Cotty, P.J., and Bandyopadhyay, R. (2014) Field efficacy of a mixture of atoxigenic Aspergillus flavus vegetative compatibility groups in preventing aflatoxin contamination in maize (Zea mays L.). Biological Control 72: 62–70.
Bandyopadhyay, R., and Cardwell, K.F. (2003) Species of Trichoderma and Aspergillus as biological control agents against plant diseases in Africa. In Biological Control in Integrated Pest Management Systems in Africa. Neuenschwander, P., Borgemeister, C., and Langewald, J. (eds). Wallingford, UK: CABI Publishing, pp. 193–206.
Bandyopadhyay, R., and Cotty, P.J. (2013) Biological Controls for aflatoxin reduction. In Aflatoxins – Finding Solutions for Improved Food Safety. Unnevehr, L., and Grace, D. (eds). Washington, DC: International Food Policy Research Institute, pp. 16–17.
Bandyopadhyay, R., Kumar, M., and Leslie, J.F. (2007) Relative severity of aflatoxin contamination of cereal crops in West Africa. Food Addit Contam 24: 1109–1114.
Barros, G.G., Torres, A.M., Rodriguez, M.I., and Chulze, S.N. (2006) Genetic diversity within Aspergillus flavus strains isolated from peanut-cropped soils in Argentina. Soil Biol Biochem 38: 145–152.
Bayman, P., and Cotty, P.J. (1991a) Improved media for selecting nitrate-non utilizing mutants in Aspergillus flavus. Mycologia 83: 311–316.
Bayman, P., and Cotty, P.J. (1991b) Vegetative compatibility and genetic diversity in the Aspergillus flavus population of a single field. Can J Bot 69: 1707–1711.
Bayman, P., and Cotty, P.J. (1993) Genetic diversity in Aspergillus flavus: association with aflatoxin production and morphology. Can J Bot 71: 23–31.
Brooker, N.L., Leslie, J.F., and Dickman, M.B. (1991) Nitrate non-utilizing mutants of Colletotrichum and their use in vegetative compatibility and genetic relatedness. Phytopathology 81: 672–677.
Cai, G., Gale, L.R., Schneider, R.W., Kistler, H.C., Davis, R.M., Elias, K.S., and Miyao, E.M. (2003) Origin of race 3 of Fusarium oxysporium f sp. lycopersici at a single site in California. Phytopathology 93: 1014–1022.
Clark, C.A., Hoy, M.W., and Nelson, P.E. (1995) Variation among isolates of Fusarium lateritium from sweet-potato for pathogenicity and vegetative compatibility. Phytopathology 85: 624–629.
Correll, J.C., Gordon, T.R., and McCain, A.H. (1988) Vegetative compatibility and pathogenicity of Verticillium alboatrum. Phytopathology 78: 1017–1021.
Cotty, P.J. (1989) Virulence and cultural characteristics of two Aspergillus flavus strains pathogenic on cotton. Phytopathology 79: 808–814.
Cotty, P.J. (1997) Aflatoxin-producing potential of communities of Aspergillus section Flavi from cotton producing areas in the United States. Mycol Res 101: 698–704.
Cotty, P.J. (2006) Biocompetitive exclusion of toxigenic fungi. In The Mycotoxin Factbook: Food and Feed Topics. Barug, D., Bhatnagar, D., van Egmond, H.P., van der Kamp, J.W., van Osenbruggen, W.A., and Visconti, A. (eds). Wageningen, The Netherlands: Wageningen Academic Publishers, pp. 179–197.
Cotty, P.J., and Bayman, P. (1993) Competitive exclusion of a toxigenic strain of Aspergillus flavus by an atoxigenic strain. Phytopathology 83: 1283–1287.
Cotty, P.J., and Cardwell, K.F. (1999) Divergence of West African and North American communities of Aspergillus section Flavi. Appl Environ Microbiol 65: 2264– 2266.
Cotty, P.J., and Sobek, E.A. (1997) The use of Aspergillus flavus to prevent aflatoxin contamination in commercial agriculture. (Abstr.) Aflatoxin elimination workshop, Memphis, Tennessee.
Cotty, P.J., Bayman, P., Egel, D.S., and Elias, K.S. (1994) Agriculture, aflatoxins and Aspergillus. In The Genus Aspergillus: From Taxonomy and Genetics to Industrial Application. Powel, K.A., Renwick, A., and Peverdy, J.F. (eds). New York, NY: Plenum Press, pp. 1–27.
Cotty, P.J., Probst, C., and Jaime-Garcia, R. (2008) Etiology and management of aflatoxin contamination. In Mycotoxins: detection methods, management, public health and agricultural trade. Leslie, J.F., Bandyopadhyay, R., and Visconti, A. (eds). Wallingford: CABI Publ, pp. 287–299.
Cove, D.J. (1976) Chlorate toxicity in Aspergillus nidulans: the selection and characterization of chlorate resistance mutants. Heredity 36: 191–203.
Cray, J.A., Bell, A.N.W., Bhaganna, P., Mswaka, A.Y., Timson, D.J., and Hallsworth, J.E. (2013) The biology of habitat dominance; can microbes behave as weeds? Microb Biotechnol 6: 453–492.
Cray, J.A., Houghton, J.D.R., Cooke, L.R., and Hallsworth, J.E. (2015) A simple inhibition coefficient for quantifying potency of biocontrol agents against plant-pathogenic fungi. Biol Cont 81: 93–100.
Donner, M., Atehnkeng, J., Sikora, R.A., Bandyopadhyay, R., and Cotty, P.J. (2009) Distribution of Aspergillus section Flavi in soils of maize fields in three agroecological zones of Nigeria. Soil Biol Biochem 41: 37–44.
Donner, M., Atehnkeng, J., Sikora, R.A., Bandyopadhyay, R., and Cotty, P.J. (2010) Molecular characterization of atoxigenic strains for biological control of aflatoxins in Nigeria. Food Addit Contam 27: 576–590.
Dorner, J.W. (2008) Development of biocontrol technology to manage aflatoxin contamination in peanuts. Peanut Sci 36: 60–67.
van Egmond, H.P., Schothorst, R.C., and Jonker, M.A. (2007) Regulations relating to mycotoxins in food: perspectives in a global and European context. Anal Bioanal Chem 389: 147–157.
Ehrlich, K.C. (2014) Non-aflatoxigenic Aspergillus flavus to prevent aflatoxin contamination in crops: advantages and limitations. Front Microbiol 5: 50. doi:10.3389/ fmicb.2014.00050.
Geiser, D.M., Frisvad, J.C., and Taylor, J.W. (1998) Evolutionary relationships in Aspergillus section Fumigati inferred from partial beta-tubulin and hydrophobin DNA sequences. Mycologia 90: 831–845.
Goss, E.M., Carbone, I., and Grünwald, N.J. (2009) Ancient isolation and independent evolution of the three clonal lineages of the exotic sudden oak death pathogen Phytophthora ramorum. Mol Ecol 18: 1161–1174.
Grace, D., Mahuku, G., Hoffmann, V., Atherstone, C., Upadhyaya, H.D., and Bandyopadhyay, R. (2015) International agricultural research to reduce food risks: case studies on aflatoxins. Food Sec 7: 569–582.
Grubisha, L., and Cotty, P.J. (2010) Genetic isolation among sympatric vegetative compatibility groups of the aflatoxinproducing fungus Aspergillus flavus. Mol Ecol 19: 269– 280.
Grubisha, L., and Cotty, P.J. (2015) Genetic analysis of the Aspergillus flavus vegetative compatibility group to which a biological control agent that limits aflatoxin contamination in USA crops belongs. Appl Environ Microbiol 81: 5889– 5899. doi:10.1128/AEM.00738-15.
Grünwald, N.J., Goodwin, S.B., Milgroom, M.G., and Fry, W.E. (2003) Analysis of genotypic diversity data for populations of microorganisms. Phytopathology 93: 738–746.
Harveson, R.M., and Rush, C.M. (1997) Genetic variation among Fusarium oxysporum isolates from sugar beet as determined by vegetative compatibility. Plant Dis 81: 85–88.
Horn, B.W. (2007) Biodiversity of Aspergillus section Flavi in the United States: a review. Food Addit Contam 24: 1088– 1101.
Horn, B.W., and Greene, R.L. (1995) Vegetative compatibility within populations of Aspergillus flavus, A. parasiticus and A. tamarii from a peanut field. Mycologia 87: 324–332.
Hruska, Z., Rajasekaran, K., Yao, H., Kincaid, R., Darlington, D., Brown, R.L., et al. (2014) Co-inoculation of aflatoxigenic and nonaflatoxigenic strains of Aspergillus flavus to study fungal invasion, colonization, and competition in maize kernels. Front Microbiol 5: 122. doi:10.3389/ fmicb.2014.00122.
Huang, C., Jha, A., Sweany, R., DeRobertis, C., and Damann, K.E., Jr (2011) Intraspecific aflatoxin inhibition is thigmoregulated, independent of vegetative compatibility group and is strain dependent. PLoS ONE 6: e23470.
Kamvar, Z.N., Tabima, J.F., and Grüwald, N.J. (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2: e281.
Keiper, F.J., Haque, M.S., Hayden, M.J., and Park, R.F. (2006) Genetic diversity in Australian populations of Puccinia graminis f. sp. avenae. Phytopathology 96: 96–104.
Leslie, J.F. (1996) Fungal vegetative compatibility – promises and prospects. Phytoparasitica 24: 3–6.
Lewis, L., Onsongo, M., Njapau, H., Schurz-Rogers, H., Luber, G., Kieszak, S., et al. (2005) Aflatoxin contamination of commercial maize products during an outbreak of acute aflatoxicosis in Eastern and Central Kenya. Environ Health Persp 113: 1763–1767.
Lewis, M.H., Carbone, I., Payne, G.A., Bowen, K.L., Hagan, A., Kemerait, R., et al. (2013) Genetic structure of soil populations of Aspergillus section Flavi and efficacy of biocontrol of aflatoxin in corn. Phytopathology 103 (Suppl. 2): S2.79.
de Lima Alves, F., Stevenson, A., Baxter, E., Gillion, J.L.M., Hejazi, F., Hayes, S., et al. (2015) Concomitant osmotic and chaotropicity-induced stresses in Aspergillus wentii: compatible solutes determine the biotic window. Curr Genet 61: 457–477.
Mauro, A., Battilani, P., Callicott, K.A., Giorni, P., Pietri, A., and Cotty, P.J. (2013) Structure of an Aspergillus flavus population from maize kernels in northern Italy. Int J Food Microbiol 162: 1–7.
Medina, A., Schmidt-Heydt, M., Rodríguez, A., Parra, R., Geisen, R., and Maga, N. (2015) Impacts of environmental stress on growth, secondary metabolite biosynthetic gene clusters and metabolite production of xerotolerant/ xerophilic fungi. Curr Genet 61: 325–334.
Mehl, H.L., and Cotty, P.J. (2010) Variation in competitive ability among isolates of Aspergillus flavus from different vegetative compatibility groups during maize infection. Phytopathology 100: 150–159.
Mehl, H.L., and Cotty, P.J. (2011) Nutrient environments influence competition among Aspergillus flavus genotypes. Appl Environ Microbiol 79: 1473–1480.
Mehl, H.L., and Cotty, P.J. (2013) Influence of plant host species on intraspecific competition during infection by Aspergillus flavus. Plant Pathol 62: 1310–1318.
Mehl, H.L., Jaime, R., Callicott, K.A., Probst, C., Garber, N.P., Ortega-Beltran, A., et al. (2012) Aspergillus flavus diversity on crops and in the environment can be exploited to reduce aflatoxin exposure and improve health. Ann N Y Acad Sci 1273: 7–17.
Moore, G.G., Elliott, J.L., Singh, R., Horn, B.W., Dorner, J.W., Stone, E.A., et al. (2013) Sexuality generates diversity in the aflatoxin gene cluster: evidence on a global scale. PLoS Pathog 9: e1003574. doi:10.1371/journal.ppat.1003574.
Novas, M.V., and Cabral, D. (2002) Association of mycotoxin and sclerotia production with compatibility groups in Aspergillus flavus from peanut in Argentina. Plant Dis 86: 215–219.
Papa, K.E. (1986) Heterokaryon incompatibility in Aspergillus flavus. Mycologia 78: 98–101.
Peraica, M., Radic´, B., Lucic´, A., and Pavlovic´, M. (1999) Toxic effects of mycotoxins in humans. Bull World Health Organ 77: 754–766.
Pildain, M.B., Vaamonde, G., and Cabral, D. (2004) Analysis of population structure of Aspergillus flavus from peanut based on vegetative compatibility, geographic origin, mycotoxin and sclerotia production. Int J Food Microbiol 93: 31–40.
Probst, C., Bandyopadhyay, R., Price, L.E., and Cotty, P.J. (2011) Identification of atoxigenic Aspergillus flavus isolates to reduce aflatoxin contamination of maize in Kenya. Plant Dis 95: 212–218.
Probst, C., Bandyopadhyay, R., and Cotty, P.J. (2014) Diversity of aflatoxin-producing fungi and their impact on food safety in sub-Saharan Africa. Int J Food Microbiol 174: 113–122.
Puhalla, J.E. (1985) Classifications of strains of Fusarium oxysporum on the basis of vegetative compatibility. Can J Bot 63: 179–183.
Schmidt, C.W. (2013) Breaking the mold – new strategies for fighting aflatoxins. Environ Health Persp 121: A270–A275.
Shannon, C.E., and Weaver, W. (1949) The Mathematical Theory of Communication. Urbana, IL: University of Illinois Press.
Shiferaw, B., Prasanna, B.M., Hellin, J., and Bänziger, M. (2011) Crops that feed the world 6. Past successes and future challenges to the role played by maize in global food security. Food Sec 3: 307–327.
Stevenson, A., Cray, J.A., Williams, J.P., Santos, R., Sahay, R., Neuenkirchen, N., et al. (2015) Is there a common water-activity limit for the three domains of life? ISME J 9: 1333–1351.
Sweany, R.R., Damann, K.E., and Kaller, M.D. (2011) Comparison of soil and corn kernel Aspergillus flavus populations: evidence for niche specialization. Phytopathology 101: 952–959.
Udoh, J.M., Cardwell, K.F., and Ikotun, T. (2000) Storage structures and aflatoxin content of maize in five agroecological zones of Nigeria. J Stored Prod Res 36: 187–201.
Varga, J., Frisvad, J.C., and Samson, R.A. (2011) Two new aflatoxin producing species, and an overview of Aspergillus section Flavi. Stud Mycol 69: 57–80.
Wicklow, D.T., and Horn, B.W. (2007) Association between vegetative compatibility and aflatoxin production by Aspergillus species during intraspecific competition. Mycoscience 48: 267–273.
Wicklow, D.T., Bobell, J.R., and Palmquist, D.E. (2003) Effect of intraspecific competition by Aspergillus flavus on aflatoxin formation in suspended disc culture. Mycol Res 107: 617–623.