Investigation of Biosynthesized Silver Nano Particles Interaction From Halymenia Porphyroides With The E7 Protein Using Bioinformatics Tool
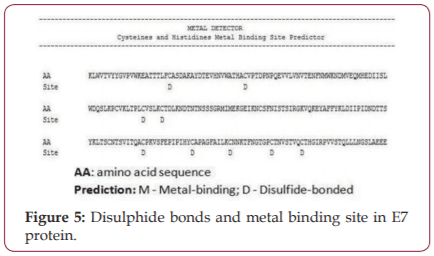
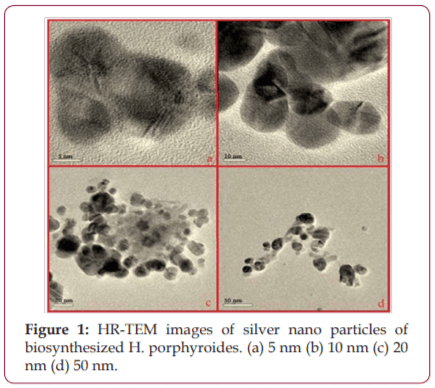
Silver nano particles have known to possess anticancer properties in the early stages of virus proliferation. In the present study biosynthesis of silver nano particles from marine red alga Halymenia porphyroides was synthesized and characterized. Studies on protein disulphide bonds were done using Metal Detector Predicts V2.0 software. All Cys and His residues in amino acid sequences were identified. The study revealed that silver nano particles couple with the sulfhydryl groups led to decrease the disulphide bonds and denatures the E7 protein thus eliminating the possibility of the occurrence of the disease.
Keywords: Bioinformatics; E7 protein; Silver nano particles; Metal binding sites.


1. Basavaraja S, Balaji SD, Lagashetty A, Rajasab AH, Venkataraman A (2008) Extracellular biosynthesis of silver nanoparticles using the fungus Fusarium semitectum. Mater Res Bull 43: 1164–1177.
2. Gardea-Torresdey JL, Gomez E, Peralta-Videa J, Parsons JG, Troiani HE (2002) Formation and growth of Au nano particles inside live alfalfa plants. Nano Lett 2: 397-401.
3. Prathna TC, Chandrasekaran NA, Raichur M, Mukherjee A (2011) Kinetic evolution study of silver nano particles in bio-based green synthesis process. Colloids Surf A Physicochem Eng 377: 212–216.
4. Rastogi L, Arunachalam J (2011) Sunlight based irradiation strategy for rapid green synthesis of highly stable silver nano particles using aqueous garlic (Allium sativum) extract and their antibacterial potential. Mater Chem Phys 129: 558-563.
5. Liu Y, Zhang YA, Zhang M (2010) Green hydrothermal synthesis and characterization of CdO2 nanoparticles. Mater Lett 64: 1779-1781.
6. Ali MD, Thajuddin N, Jeganathan K, Gunasekaran M (2011) Plant extract mediated synthesis of silver and gold nano particles and its antibacterial activity against clinically isolated pathogens. Colloids Surf B 85: 360- 365.
7. Vidhu VK, Aromal SA, Philip D (2011) Green synthesis of silver nanoparticles using Macrotyloma uniflorum. Spectrochim Acta A 83: 392-397.
8. Zhan G, Huang J, Du M, Abdul-Rauf I, Ma Y, et al. (2011) Green synthesis of Au–Pd bimetallic nano particles: Single-step bioreduction method with plant extract. Mater Lett 65: 2989-2991.
9. Dubey SP, Lahtinen M, Sillanpää M (2010) Tansy fruit mediated greener synthesis of silver and gold nanoparticles. Process Biochem 45: 1065- 1071.
10. Kumar VG, Gokavarapu SD, Rajeswari A, Dhas TS, Karthick V, et al. (2011) Facile green synthesis of gold nanoparticles using leaf extract of antidiabetic potent Cassia auriculata. Colloids Surf B 87: 159-163.
11. Smitha SL, Philip D, Gopchandran KG (2009) Green synthesis of gold nanoparticles using Cinnamomum zeylanicum leaf broth. Spectrochim. Acta A 74: 735-739.
12. Dwivedi A, Gopal K (2010) Biosynthesis of silver and gold nano particles using Chenopodium album leaf extract. Colloid Surface A: Physicochem Eng Asp 369: 27-33.
13. Li Y, Wu Y, Ong BS (2005) Facile synthesis of silver nano particles useful for fabrication of high-conductivity elements for printed electronics. J Am Chem Soc 127: 3266-3267.
14. Evanoff DD, Chumanov G (2005) Synthesis and optical properties of silver nano particles and arrays. Chem Phys Chem 6: 1221-1231.
15. Sun Y, Xia Y (2002) Shape-controlled synthesis of gold and silver nano particles. Science 298: 2176-2179.
16. Sun RWY, Chen R, Chung NPY, Ho CM, Lin CLS, et al. (2005) Silver nano particles fabricated in Hepes buffer exhibit cytoprotective activities toward HIV-1 infected cells. Chem Com 40: 5059-5061.
17. Lu L, Sun R, Chen R, Hui CK, Ho CM, et al. (2008) Silver nano particles inhibit hepatitis B virus replication. Antiviral Therapy 13(2): 253-262.
18. Sun L, Singh AK, Vig K, Pillai SR, Singh SR (2008) Silver nano particles inhibit replication of respiratory syncytial virus. J Biomed Nanotech 4(2): 149-158.
19. Baram-Pinto D, Shukla S, Perkas N, Gedanken A, Sarid R (2009) Inhibition of herpes simplex virus type 1 infection by silver nano particles capped with mercaptoethane sulfonate. Bioconjugate Chemistry 20(8): 1497- 1502.
20. Rogers JV, Parkinson CV, Choi YW, Speshock JL, Hussain SM (2008) A preliminary assessment of silver nano particle inhibition of monkey pox virus plaque formation. Nanoscale Research Letters 3(4): 129-133.
21. KE Clemens, R Brent, J Gyuris, K Munger (1995) Dimerization of the human papilloma virus E7 oncoprotein in vivo. Virology 214: 289-293.
22. CC Baker, WC Phelps, Lindgren V, MJ Braun, MA Gonda, et al. (1987) Structural and transcriptional analysis of human papilloma virus type 16 sequences in cervical carcinoma cell lines. Journal of Virology 61(4): 962-971.
23. K Munger, WC Phelps (1993) The human papilloma virus E7 protein as a transforming and transactivating factor. Biochimica et Biophysica Acta 1155(1): 111-123.
24. Vishnu Kiran M, Murugesan S (2014) Biosynthesis of silver nano particles from marine alga Halymenia poryphyroides and its antibacterial efficacy. Int J Curr Microbiol App Sci 3(4): 96-103.
25. Vishnu Kiran M, Murugesan S (2014) Biological synthesis of silver nano particles from marine alga Colpomenia sinuosa and its in vitro antidiabetic activity. AJBBL 3(1): 1-7.
26. Vishnu Kiran M, Murugesan S (2013) Biogenic silver nanoparticles by Halymenia porphyroides and its in vitro anti-diabetic efficacy. J Chem Pharma Res 5(12): 1001-1008.
27. Barbosa MS, Edmonds C, Fisher C, Schiller JT, Lowy DR et al. (1990) The region of the hpv E7 oncoprotein homologous to adenovirus E1a and Sv40 large T antigen contains separate domains for Rb binding and casein kinase II phosphorylation. Embo Journal 9(1): 153-160.
28. Lara HH, Ayala-Nunez NV, Ixtepan-Turrent L, Rodriguez-Padilla C (2010) Mode of antiviral action of silver nano particles against HIV-1. J Nanobiotechnol 8: 1-10.
29. Shahin Gavanji, Hassan Mohabatkar, Hojjat Baghshahi, Ali Zarrabi (2014) Bioinformatics Prediction of Interaction Silver Nano particles on the Disulfide Bonds of HIV-1 Gp120 Protein. International Journal of Scientific Research in Knowledge 2(2): 67-74.



.jpg&w=3840&q=75)